Which files do I need to upload for Spectronaut DIA processed results?
Mass Dynamics supports both protein-only and peptide + protein analyses for Spectronaut DIA data. To simplify your upload process and avoid working with the large BGS Normal Report, we provide lightweight Mass Dynamics export schemas. These schemas contain only the minimum required columns needed for analysis, producing smaller files that are faster to upload and process.
You can choose between:
- Option 1: Peptide + Protein Mass Dynamics Schema - Download here [Recommended]
- Option 2: Protein-Only Mass Dynamics Schema - Download here
- Option 3: BGS Normal Report [Not Recommended]
Below are the details and steps for each option.
OPTION 1 - Peptide + Protein Mass Dynamics Schema [Recommended]
This Mass Dynamics schema allows you to generate a smaller report containing the minimum required columns to perform protein and peptide analysis in Mass Dynamics.
Step to generate the required processed output from Spectronaut:
- Download the Peptide+Protein Mass Dynamics Schema here.
- Import the schema to Spectronaut.
- Setup the appropriate Grouping Requirements in Spectornaut (see next section)
- Export the Spectronaut Report.tsv
- Go to the Upload Page in Mass Dynamics, upload the Report.tsv.
Grouping Requirements
Peptide-level information, including the modified sequence and its intensity, is derived from the columns GroupingKey, GroupingKeyType, and PEP.Quantity.
If you expect to observe post-translational modifications (PTMs) in your data, selecting the modified sequence option is strongly recommended; otherwise, modification-specific information will not be available in Mass Dynamics.
For analysis at the modified sequence level:
- Select “Modified Sequence” in the quantification grouping within the analysis settings (see screenshot below).
- This ensures that PEP.Quantity in the exported report reflects the intensities of modified sequences, and GroupingKey corresponds to the modified sequence.
If instead “Stripped Sequence” is selected:
- Both PEP.Quantity and GroupingKey will correspond to the unmodified (stripped) sequence. As a result, PTM analysis will not be possible.
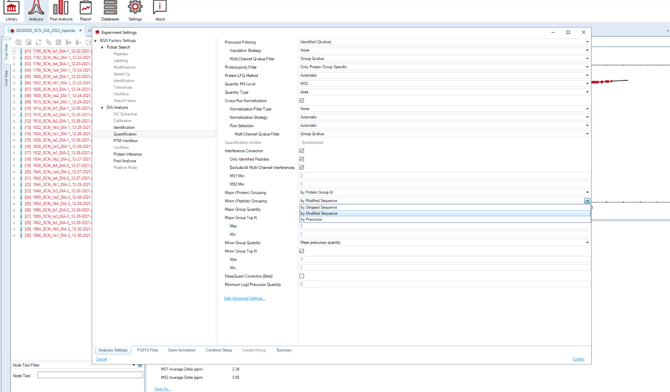
Spectronaut grouping requirement for peptide-level analyses.
OPTION 2 - Protein-Only Mass Dynamics Schema
This schema contains the minimal required columns to perform protein-only analysis in Mass Dynamics.
Step to generate the required processed output from Spectronaut:
- Download the Protein-Only Mass Dynamics Schema here.
- Import the Schema to Spectronaut.
- Export the Spectronaut Report.tsv
- Go to the Upload Page in Mass Dynamics, upload the Report.tsv.
OPTION 3 - BGS Normal Report [Not Recommended]
The BGS Normal Report must be generated with the following extra options enabled in the settings: “PG.Genes”, “PEP.Quantity” (see screenshot below; “PG.Description” is optional).
The peptide intensities will be extracted from “PEP.Quantity” which is not by default included in the BGS Normal report.
Step to generate the required processed output from Spectronaut:
- Export the Spectronaut BGS Report.tsv
- Go to the Upload Page in Mass Dynamics, upload the Report.tsv.
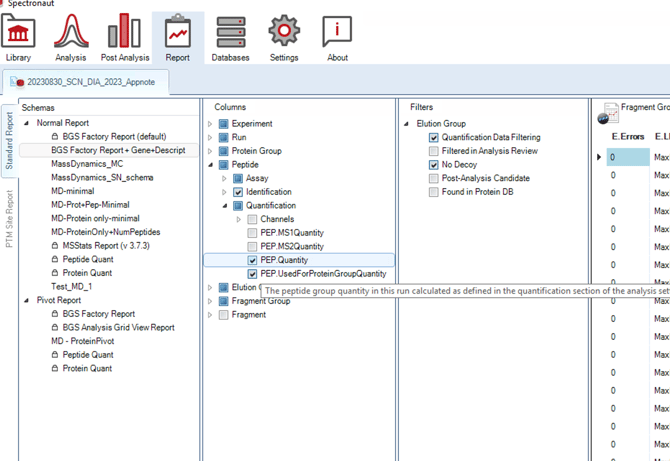
Column requirements to enable peptide-level analysis with the BGS Normal Report
If you need support uploading your data, you can follow these instructions to leverage our assisted upload service.